Module 3: MHC/DLA Typing
To trigger an immune response, a mutant protein fragment (neoantigen) must be physically presented on the tumor cell surface by MHC molecules. In dogs, the MHC system is called DLA (Dog Leukocyte Antigen). This module determines which DLA alleles the patient carries and confirms they are actively expressed, because only those specific alleles will determine which neoantigens can actually be presented.
Prepare DLA Reference
The DLA system consists of highly polymorphic genes that encode the proteins responsible for presenting peptide fragments to immune cells. Different alleles present different sets of peptides, so knowing the exact DLA type is critical for predicting which neoantigens will be visible to the immune system.
We download the complete set of MHC nucleotide sequences from the IPD-MHC database[1], the authoritative repository for MHC sequences across species. From this collection, we extract only the canine DLA alleles, rename the headers to a clean format (e.g., DLA-88*001:01), and deduplicate by keeping the longest sequence per allele name.
Two separate indices are built for different purposes: a BWA index for aligning WES DNA reads in Step 02, and a Salmon index[2] for quantifying RNA-seq expression in Step 03.
# Download MHC nucleotide sequences from IPD-MHC
wget -O MHC_nuc.fasta \
https://raw.githubusercontent.com/ANHIG/IPDMHC/Latest/MHC_nuc.fasta
# Extract DLA sequences and rename headers
awk '/^>/{if($2 ~ /^DLA-/) {print ">"$2; keep=1} else {keep=0}; next} keep{print}' \
MHC_nuc.fasta > DLA_nuc.raw.fasta
# Deduplicate: keep longest sequence per allele name
# (omitted for brevity)
# Build BWA index (for WES typing)
bwa index DLA_nuc.fasta
# Build Salmon index (for RNA-seq quantification)
salmon index -t DLA_nuc.fasta -i salmon_indexWES-based DLA Typing
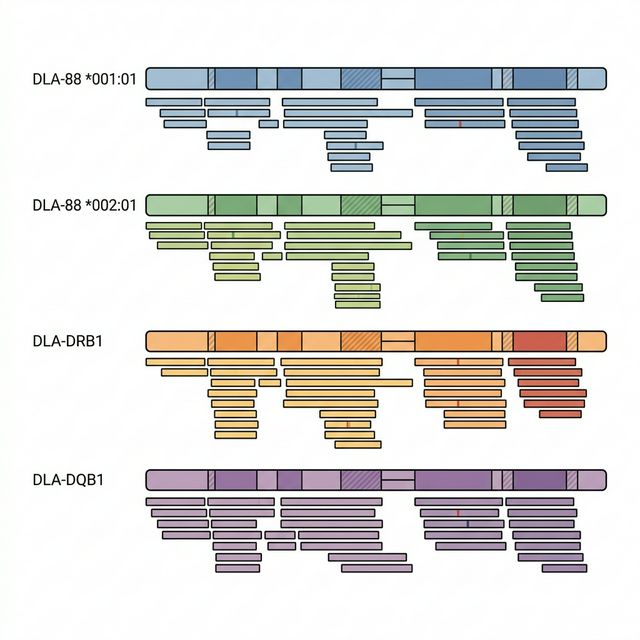
DLA typing uses the normal (germline) WES data from Module 1, not the tumor data. This is because DLA alleles are inherited and constant across all cells in the body. Using normal tissue avoids any confounding mutations or copy-number changes that might distort the signal in tumor DNA.
The trimmed WES reads are aligned to the DLA reference using BWA-MEM, and only mapped reads are retained. Then, for each DLA allele, we calculate two key metrics using samtools coverage: coverage breadth (what fraction of the allele sequence is covered by at least one read) and mean depth (average number of reads per base position).
The typing algorithm works as follows: for each of the seven DLA loci (DLA-88, DLA-12, DLA-64, DLA-79, DLA-DRB1, DLA-DQA1, DLA-DQB1), we select all alleles with coverage breadth ≥ 50% as candidates. Alleles are ranked by coverage breadth first, then by mean depth as a tiebreaker. This threshold-based approach is more robust than simply picking the top 2, as it naturally handles homozygous loci (where only one allele should be selected) and avoids forcing a second allele when no confident match exists.
# Align normal WES reads to DLA reference
bwa mem -t $THREADS $DLA_REF \
trimmed/${SRR}_1.trimmed.fastq.gz \
trimmed/${SRR}_2.trimmed.fastq.gz | \
samtools view -b -F 4 - | \
samtools sort -@ $THREADS -o ${SRR}.dla.sorted.bam
samtools index ${SRR}.dla.sorted.bam
# Calculate per-allele coverage statistics
samtools coverage ${SRR}.dla.sorted.bam | \
awk -F'\t' 'NR>1 && $4>0 {print $1, $3, $4, $5, $6/100, $7}' | \
sort -k5 -nr -k6 -nr > allele_coverage.tsv.tmp
echo -e "ALLELE\tLENGTH\tMAPPED_READS\tCOVERED_BASES\tCOVERAGE_BREADTH\tMEAN_DEPTH" > allele_coverage.tsv
cat allele_coverage.tsv.tmp >> allele_coverage.tsv
# Select DLA candidates: alleles with coverage breadth >= 0.5
MIN_BREADTH=0.5
for LOCUS in DLA-88 DLA-12 DLA-64 DLA-79 DLA-DRB1 DLA-DQA1 DLA-DQB1; do
grep "${LOCUS}" allele_coverage.tsv | \
sort -k5 -nr -k6 -nr | \
awk -v min=$MIN_BREADTH '$5 >= min'
done > dla_typing_result.tsvRNA-seq DLA Expression
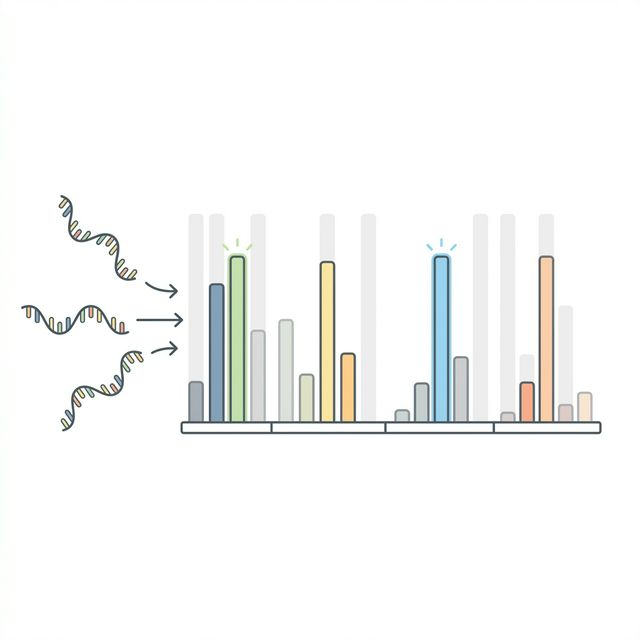
Knowing which DLA alleles exist in the genome is necessary but not sufficient. Tumors can silence MHC genes through epigenetic mechanisms or chromosomal deletions, a well-known immune evasion strategy. If a DLA allele is not expressed, it cannot present neoantigens on the cell surface, and any peptide predicted to bind that allele would be useless as a vaccine target.
We use Salmon[2], a fast quasi-mapping tool, to quantify the expression of each DLA allele directly from the tumor RNA-seq reads (same trimmed FASTQs from Module 2). Salmon uses lightweight alignment (--validateMappings) and expectation-maximization to estimate allele-level TPM.
For each DLA allele identified in Step 02, we check its expression in the Salmon output using two thresholds: TPM ≥ 5 and NumReads ≥ 10. Both conditions must be met for an allele to be classified as expressed. The read count threshold prevents low-confidence calls from a handful of stray reads, while TPM normalizes for transcript length and sequencing depth.
Finally, the pipeline produces a final DLA genotype that combines WES coverage evidence with RNA-seq expression. Only alleles that pass both the WES typing and RNA expression filters are included. This merged result is what Module 4 uses for neoantigen binding prediction.
# Quantify DLA allele expression with Salmon
salmon quant \
-i $DLA_IDX \
-l A \
-1 trimmed/${SRR}_1.trimmed.fastq.gz \
-2 trimmed/${SRR}_2.trimmed.fastq.gz \
-p $THREADS \
--validateMappings \
-o expression/${SRR}
# Check expression for each typed DLA allele
TPM_THRESHOLD=5
MIN_READS=10
# Expressed = YES if TPM >= 5 AND NumReads >= 10
# Output: LOCUS ALLELE TPM NUM_READS EXPRESSED(YES/NO)
# Produce final genotype: WES coverage + RNA expression (expressed alleles only)
# Output: LOCUS ALLELE WES_BREADTH WES_DEPTH RNA_TPM RNA_READSExpected Metrics & QC Checkpoints
Before proceeding to neoantigen prediction, verify these quality metrics from Module 3. The accuracy of DLA typing directly affects which neoantigens are predicted to bind, so false alleles will cascade into incorrect vaccine candidates.
1. DLA Alignment
- Mapped Reads> 0 per allele
- Top Allele Breadth> 80%
- Top Allele Depth> 5x
Low coverage may indicate the WES capture kit does not include MHC regions.
2. DLA Typing
- Alleles per Locus1-2
- Loci Typed7 (class I + II)
- Breadth Threshold≥ 50%
Ambiguous typing (many alleles with similar coverage) suggests homozygosity or cross-mapping.
3. DLA Expression
- TPM Threshold≥ 5
- Min Reads≥ 10
- Class I ExpressedExpected
- Class II ExpressedVariable
Absent class I expression may indicate MHC downregulation, a common immune evasion mechanism.
References
- Maccari, G. et al. (2017). IPD-MHC 2.0: an improved inter-species database for the study of the major histocompatibility complex. Nucleic Acids Research, 45(D1), D860-D864. doi:10.1093/nar/gkw1050 [IPD-MHC database]
- Patro, R. et al. (2017). Salmon provides fast and bias-aware quantification of transcript expression. Nature Methods, 14, 417-419. doi:10.1038/nmeth.4197 [Salmon]
- Li, H. (2013). Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. arXiv, 1303.3997. doi:10.48550/arXiv.1303.3997 [BWA-MEM]