Module 5: Vaccine Cassette Design & Assembly
The culmination of the pipeline is translating raw predictions into a tangible, administrable vaccine. This module takes the prioritized neoantigens from Module 4, logically strings them together into a structurally stable multi-epitope 'cassette,' and reverse-translates the protein back into a codon-optimized DNA/mRNA sequence ready for clinical manufacturing.
Epitope Selection
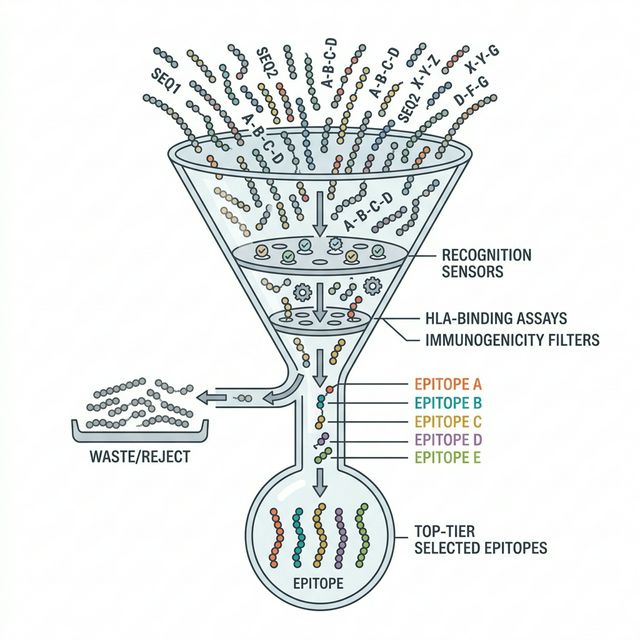
Think of the vaccine cassette as a carefully curated "greatest hits" album. We cannot include every mutated peptide, so we must select the very best targets. We filter the ranked candidates from Module 4 to select the top 10 Class I (CD8+ targets) and the top 5 Class II (CD4+ targets) epitopes.
The selection process revolves around three key principles: First, the epitope must be RNA-VALIDATED, meaning we have concrete proof it is actively being transcribed by the tumor. Second, it must have a high EL_SCOREensuring it will actually bind to the patient's MHC receptors. Finally, we deduplicate by peptide sequence to ensure we aren't wasting space on identical peptides stemming from different transcripts of the same gene.
If we lack enough fully validated targets, the script automatically backfills the remaining slots with candidates marked as WEAK_SUPPORT (expressed, but below strict depth thresholds), ensuring we always maximize the capacity of the vaccine platform.
# Select top epitopes from Class I and Class II candidates
# Class I: deduplicate by peptide, keep best EL_SCORE, prefer VALIDATED
tail -n +2 class1_epitope_candidates.tsv | sort -t$'\t' -k8 -nr | \
awk -F'\t' '!seen[$5]++ && $11=="VALIDATED"' | head -n 10
# Class II: same strategy
tail -n +2 class2_epitope_candidates.tsv | sort -t$'\t' -k8 -nr | \
awk -F'\t' '!seen[$5]++ && $11=="VALIDATED"' | head -n 5Cassette Assembly
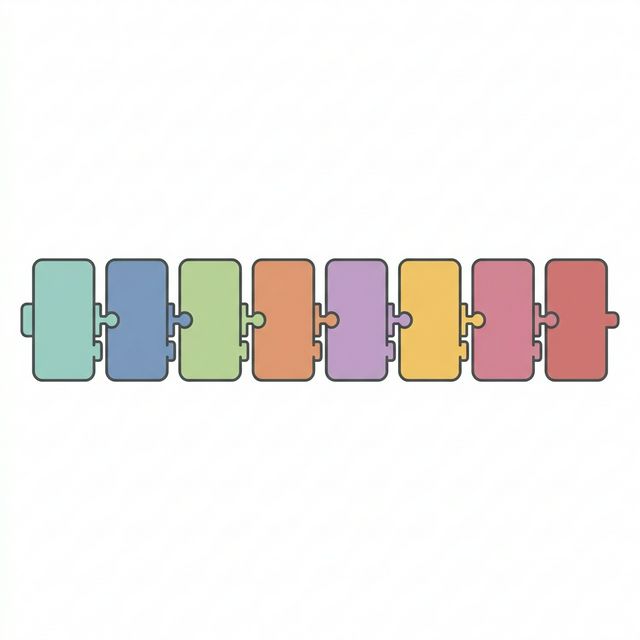

Selected epitopes cannot simply be glued together randomly. If we do, the boundaries between two adjacent epitopes might accidentally create a brand new, highly immunogenic peptide that doesn't exist in the tumor (a "junctional epitope"). Instead, we assemble them using a meticulously engineered multi-epitope architecture.
The cassette begins with a tissue plasminogen activator (tPA) signal sequence to ensure the synthesized protein is properly routed to the endoplasmic reticulum and secreted for immune recognition. Class I epitopes are separated by rigid AAY linkers, which serve as highly efficient cleavage sites for the proteasome. Class II epitopes are separated by highly flexible GPGPG linkers, minimizing structural interference. Finally, we cap the C-terminus with PADRE[2], a synthetic "universal" CD4+ helper epitope that boosts the immune response regardless of the patient's specific MHC typing.
3D Structural Verification
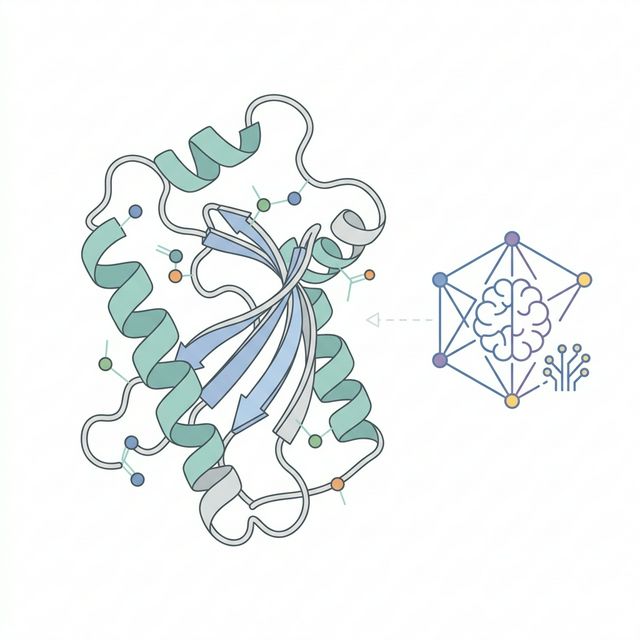
Once we have our linear amino acid sequence (typically around 250 aa), we need to ensure the final artificial protein is physically viable. We submit the sequence to ESMFold[1], Meta AI's cutting-edge language model for protein folding, running locally on a dedicated GPU (RTX 3090).
ESMFold outputs a complete set of 3D atomic coordinates (a PDB file) and per-residue confidence scores known as pLDDT. For natural proteins, a high pLDDT is crucial. However, our vaccine cassette is an artificial chimera designed specifically to be chopped up by the proteasome, not to fold into a stable globe. As such, a low pLDDT is actually expected and perfectly acceptable here.
Instead, our primary quality checkpoint is the Instability Indexcalculated via Biopython's ProtParam. If this score is below 40, we can be confident the protein will be stable enough to be synthesized and handled in vitro without rapidly degrading.
# Run ESMFold on GPU (RTX 3090)
# Model: facebook/esmfold_v1 (~8.4GB)
# Input: cassette_protein.fasta (254 aa)
mamba run -n wecanvax2 python modules/module5/03-structural_verification.py
# Output:
# cassette_3D.pdb — 3D atomic coordinates
# structural_report.json — biophysical + pLDDT summaryCodon-Optimized Reverse Translation
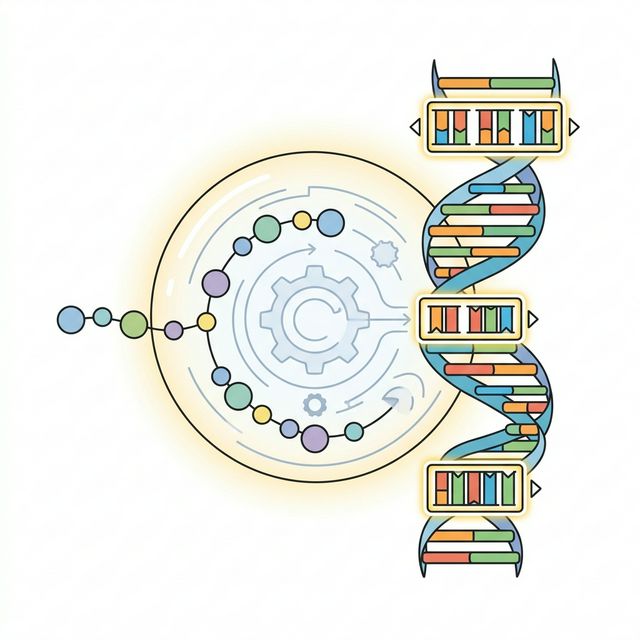

The final step before manufacturing is converting the protein back into DNA (or mRNA). Because the genetic code is redundant, there are many ways to spell the same protein. However, for mRNA vaccines, this spelling drastically impacts clinical efficacy. We use a sophisticated GC-max / U-min strategy.
First, high GC content (Guanine-Cytosine)[3]severely tightens the mRNA's thermal stability and secondary structure, dramatically increasing the half-life of the molecule inside the cell. Our algorithm parses the Canis lupus familiaris codon usage table and always prioritizes codons with the maximum GC percentage.
Second, we strictly minimize Uridine (U, or T in DNA)content. Uridine-rich tracts are notorious for triggering the innate immune system's Toll-like receptors (TLR7/8), causing the body to destroy the mRNA before it can be effectively translated into our vaccine antigens. When GC content is tied, the script strictly breaks ties in favor of the codon with fewer U/T bases.
Finally, the script automatically appends a definitive STOP codon (TGA) to prevent ribosomal readthrough into the poly-A tail, completing the pipeline.
# GC-max / U-min codon selection strategy
# For each amino acid:
# 1. Sort codons by GC content (descending)
# 2. Among ties, pick fewest T (U) bases
# 3. Append TGA stop codon
# Example — Leucine (L):
# CTG (GC=67%, T=0) ← SELECTED (highest GC, lowest U)
# CTC (GC=67%, T=1)
# CTT (GC=33%, T=2)
# TTG (GC=33%, T=2) ← NEVER usedExpected Metrics & QC Checkpoints
The completed vaccine cassette has been validated at every level: epitope selection, structural integrity, biophysical properties, and codon optimization. Review these final metrics:
1. Epitope Content
- Class I Targets10
- Class II Targets5
- RNA Validation100% PASS
Ensures we target both CD8+ cytotoxic and CD4+ helper T cell subsets.
2. Protein Biophysics
- Protein Length~250 aa
- Instability Index< 40.0
- GRAVY Score< 0 (Hydrophilic)
An instability index under 40 confirms the artificial construct is structurally viable.
3. DNA/mRNA Optimization
- GC Content≥ 70.0%
- T(U) Content≤ 15.0%
- Stop CodonTGA appended
Extreme GC maximization and U-minimization prevents unwanted RNA degradation in vivo.
References
- Lin, Z. et al. (2023). Evolutionary-scale prediction of atomic-level protein structure with a language model. Science, 379(6637), 1123–1130. doi:10.1126/science.ade2574 [ESMFold]
- Alexander, J. et al. (1994). Development of high potency universal DR-restricted helper epitopes by modification of high affinity DR-blocking peptides. Immunity, 1(9), 751–761. doi:10.1016/S1074-7613(94)80017-0 [PADRE universal helper epitope]
- Kudla, G. et al. (2006). High guanine and cytosine content increases mRNA levels in mammalian cells. PLoS Biology, 4(6), e180. doi:10.1371/journal.pbio.0040180 [GC content & expression]